Visualize a causal graph stored within a Disco object. This function
extends plot.Knowledge() by combining the causal graph from a caugi object with
background knowledge.
Usage
# S3 method for class 'Disco'
plot(x, required_col = "blue", ...)Arguments
- x
A
Discoobject containing both the causal graph and the associated knowledge.- required_col
Character(1). Color for edges marked as "required". Default
"blue".- ...
Additional arguments passed to
caugi::plot()andplot.Knowledge().
Details
Required edges are drawn in blue by default (
required_col), can be changed.Forbidden edges are not drawn.
If tiered knowledge is provided, nodes are arranged according to their tiers.
Other edge styling (line width, arrow size, etc.) can be supplied via
edge_style. To override the color of a specific edge, specify it inedge_style$by_edge[[from]][[to]]$col.
This function combines the causal graph and the Knowledge object into a single plotting
structure. If the knowledge contains tiers, nodes are laid out accordingly; otherwise,
the default caugi layout is used. Edges marked as required are automatically colored
(or can be overridden per edge using edge_style$by_edge).
Examples
data(tpc_example)
# Define tiered knowledge
kn <- knowledge(
tpc_example,
tier(
child ~ starts_with("child"),
youth ~ starts_with("youth"),
old ~ starts_with("old")
)
)
# Fit a causal discovery model
cd_tges <- tges(engine = "causalDisco", score = "tbic")
disco_cd_tges <- disco(data = tpc_example, method = cd_tges, knowledge = kn)
# Plot with default column orientation
plot(disco_cd_tges)
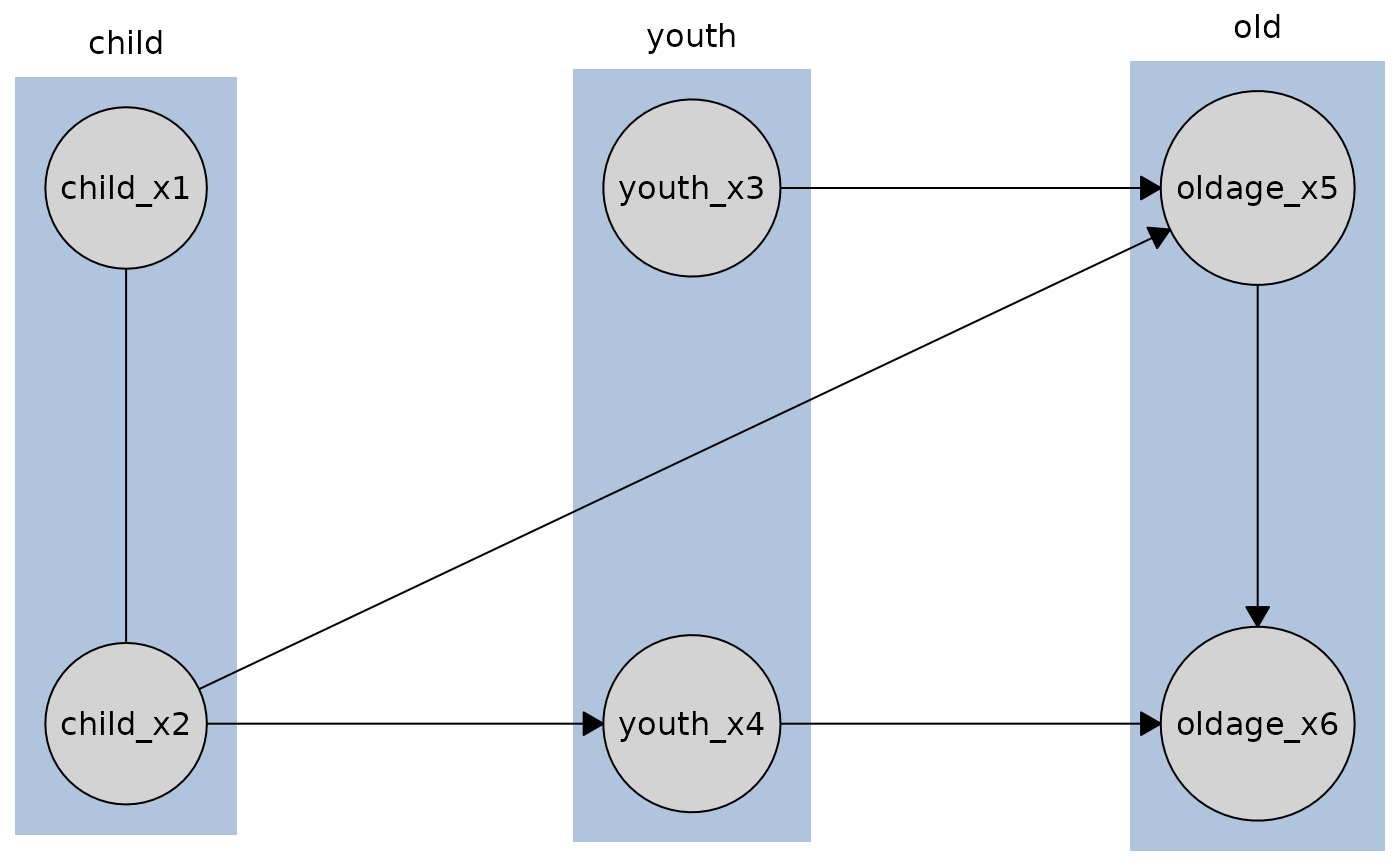 # Plot with row orientation
plot(disco_cd_tges, orientation = "rows")
# Plot with row orientation
plot(disco_cd_tges, orientation = "rows")
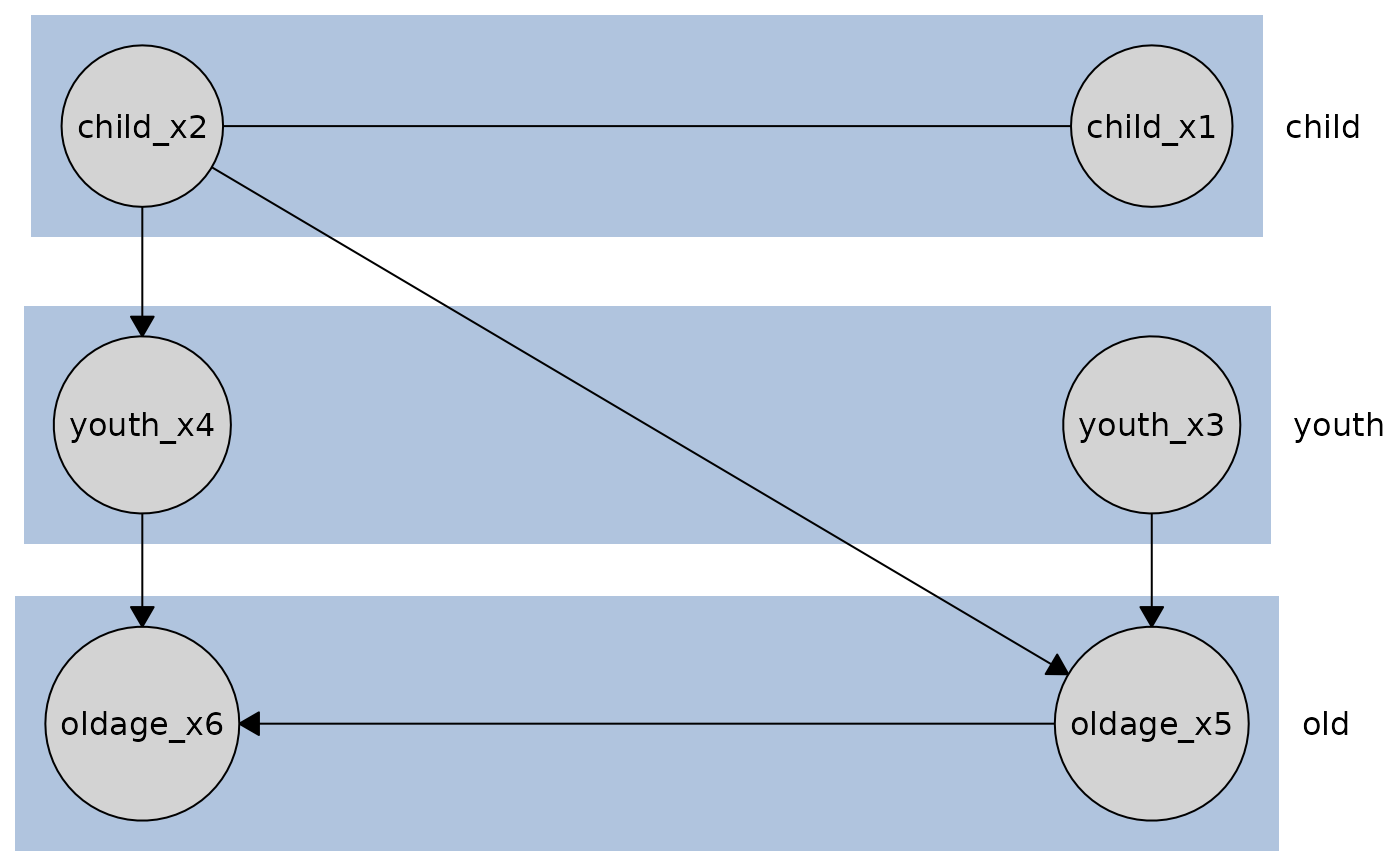 # Plot with custom node and edge styling
plot(
disco_cd_tges,
node_style = list(
fill = "lightblue", # Fill color
col = "darkblue", # Border color
lwd = 2, # Border width
padding = 4, # Text padding (mm)
size = 1.2 # Size multiplier
),
edge_style = list(
lwd = 1.5, # Edge width
arrow_size = 4, # Arrow size (mm)
col = "darkgreen", # Edge color
fill = "black", # Arrow fill color
lty = "dashed" # Edge line type
)
)
# Plot with custom node and edge styling
plot(
disco_cd_tges,
node_style = list(
fill = "lightblue", # Fill color
col = "darkblue", # Border color
lwd = 2, # Border width
padding = 4, # Text padding (mm)
size = 1.2 # Size multiplier
),
edge_style = list(
lwd = 1.5, # Edge width
arrow_size = 4, # Arrow size (mm)
col = "darkgreen", # Edge color
fill = "black", # Arrow fill color
lty = "dashed" # Edge line type
)
)
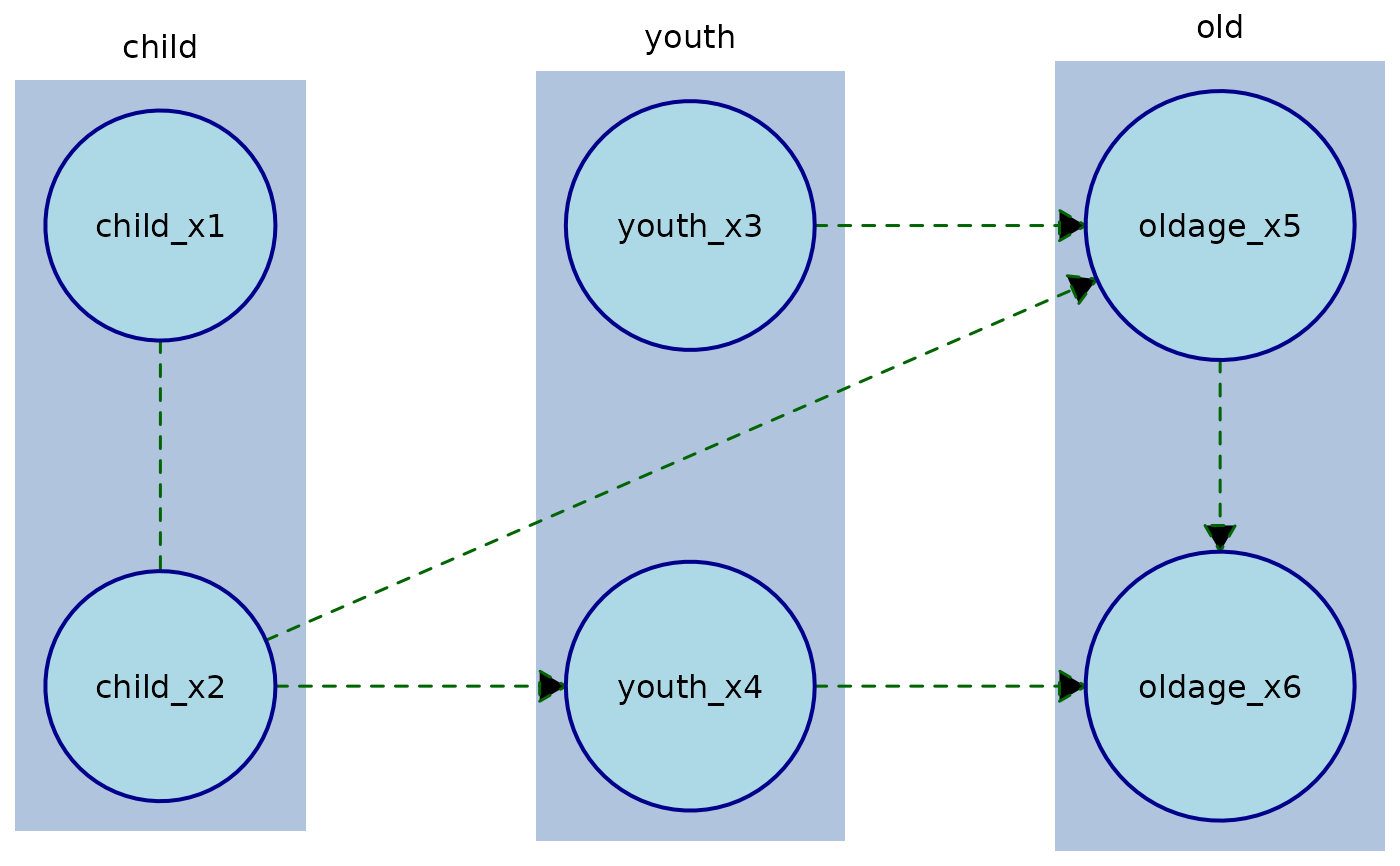 # To override a specific edge style which is required you need to target that individual node:
kn <- knowledge(
tpc_example,
tier(
child ~ starts_with("child"),
youth ~ starts_with("youth"),
old ~ starts_with("old")
),
child_x1 %-->% c(child_x2, youth_x4) # required edges
)
bnlearn_pc <- pc(engine = "bnlearn", test = "fisher_z")
disco_bnlearn_pc <- disco(data = tpc_example, method = bnlearn_pc, knowledge = kn)
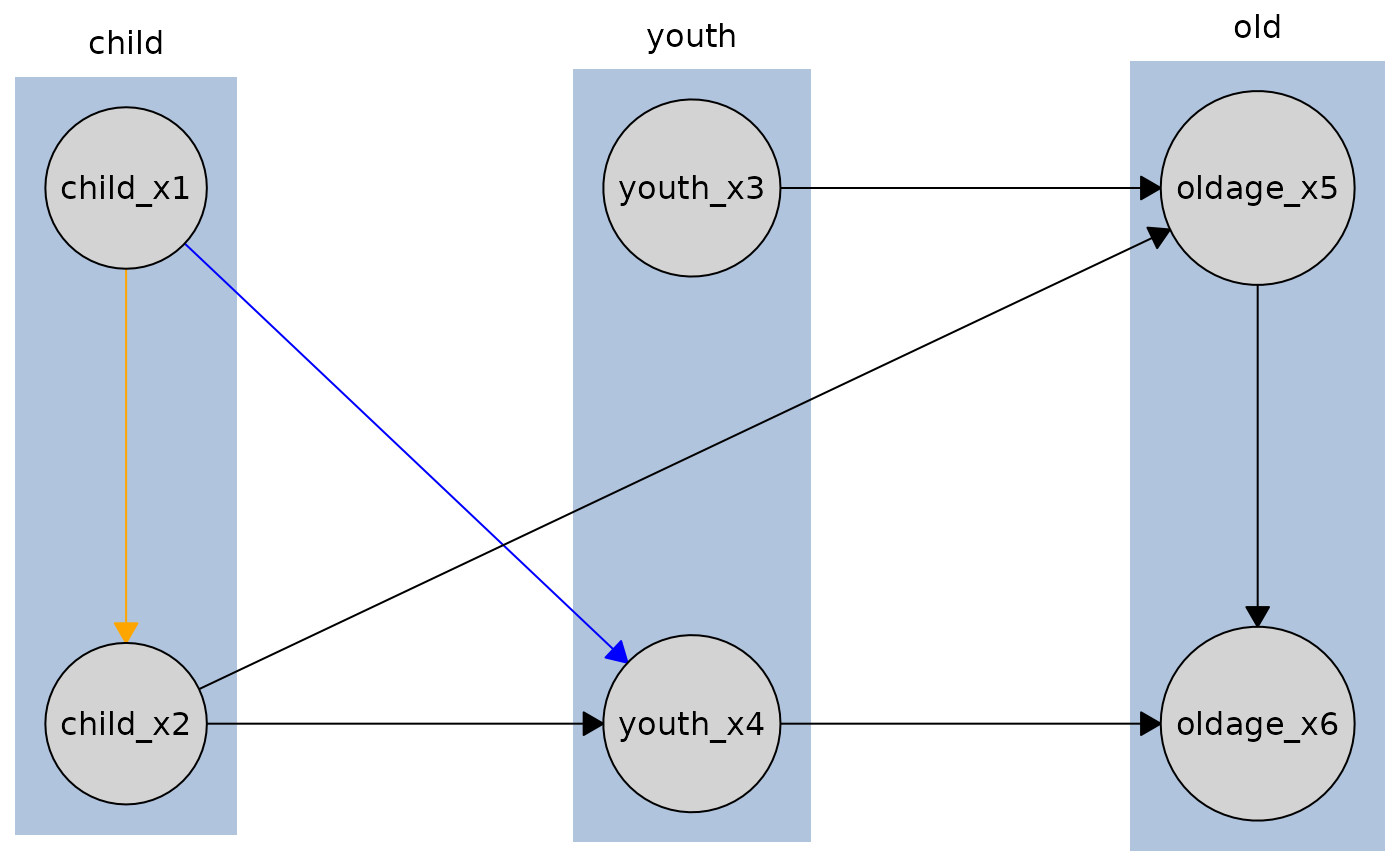
# Edge from child_x1 to child_x2 will be orange, but edge from child_x1 to youth_x4
# will be required_col (blue) since we only override the child_x1 to child_x2 edge.
plot(
disco_bnlearn_pc,
edge_style = list(
by_edge = list(
child_x1 = list(
child_x2 = list(col = "orange", fill = "orange")
)
)
),
required_col = "blue"
)
# To override a specific edge style which is required you need to target that individual node:
kn <- knowledge(
tpc_example,
tier(
child ~ starts_with("child"),
youth ~ starts_with("youth"),
old ~ starts_with("old")
),
child_x1 %-->% c(child_x2, youth_x4) # required edges
)
bnlearn_pc <- pc(engine = "bnlearn", test = "fisher_z")
disco_bnlearn_pc <- disco(data = tpc_example, method = bnlearn_pc, knowledge = kn)
# Edge from child_x1 to child_x2 will be orange, but edge from child_x1 to youth_x4
# will be required_col (blue) since we only override the child_x1 to child_x2 edge.
plot(
disco_bnlearn_pc,
edge_style = list(
by_edge = list(
child_x1 = list(
child_x2 = list(col = "orange", fill = "orange")
)
)
),
required_col = "blue"
)
 # Plot without tiers
data(num_data)
kn_untiered <- knowledge(
num_data,
X1 %-->% c(X2, X3),
Z %!-->% Y
)
bnlearn_pc <- pc(engine = "bnlearn", test = "fisher_z")
res_untiered <- disco(data = num_data, method = bnlearn_pc, knowledge = kn_untiered)
plot(res_untiered)
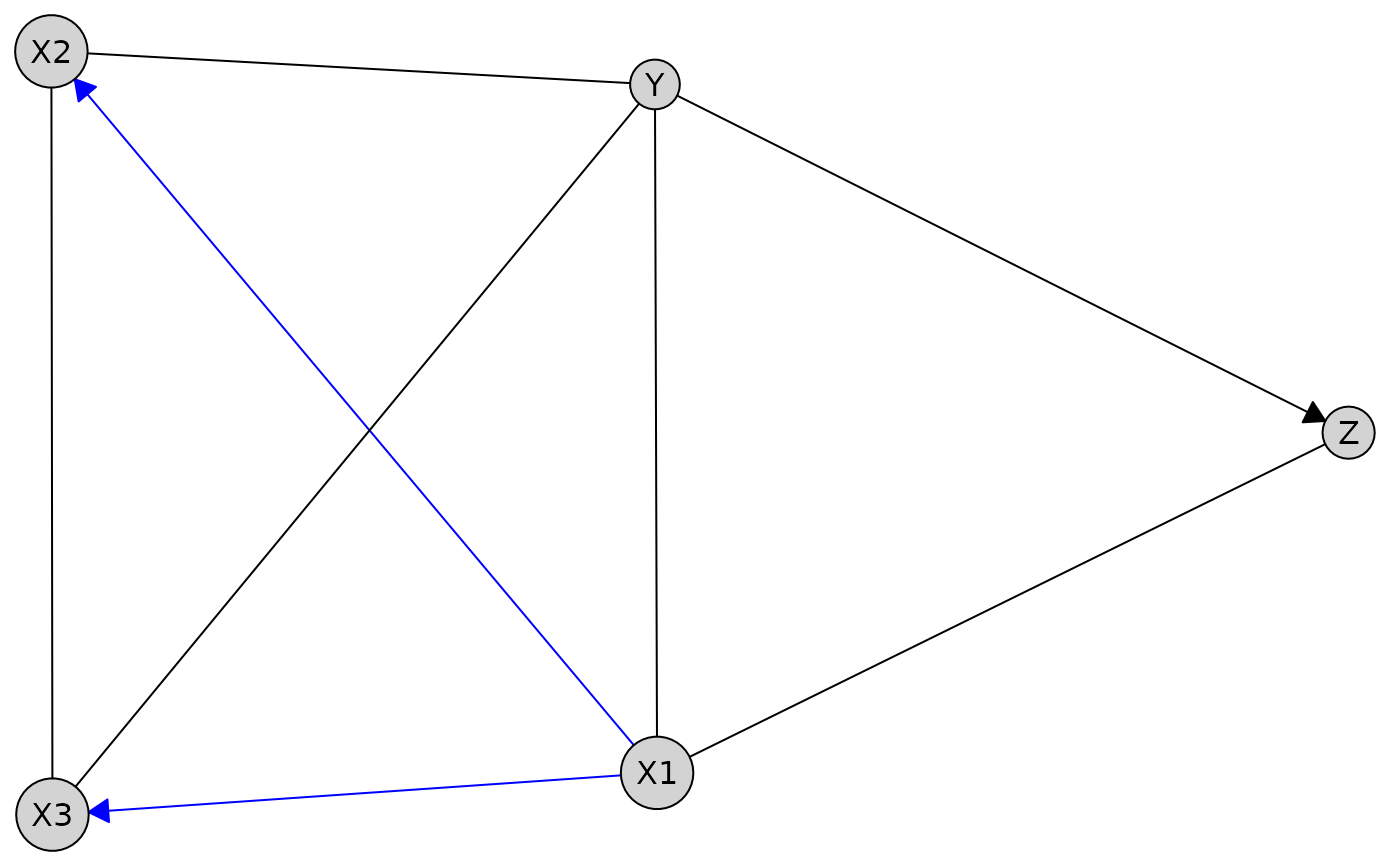
# Plot without tiers
data(num_data)
kn_untiered <- knowledge(
num_data,
X1 %-->% c(X2, X3),
Z %!-->% Y
)
bnlearn_pc <- pc(engine = "bnlearn", test = "fisher_z")
res_untiered <- disco(data = num_data, method = bnlearn_pc, knowledge = kn_untiered)
plot(res_untiered)
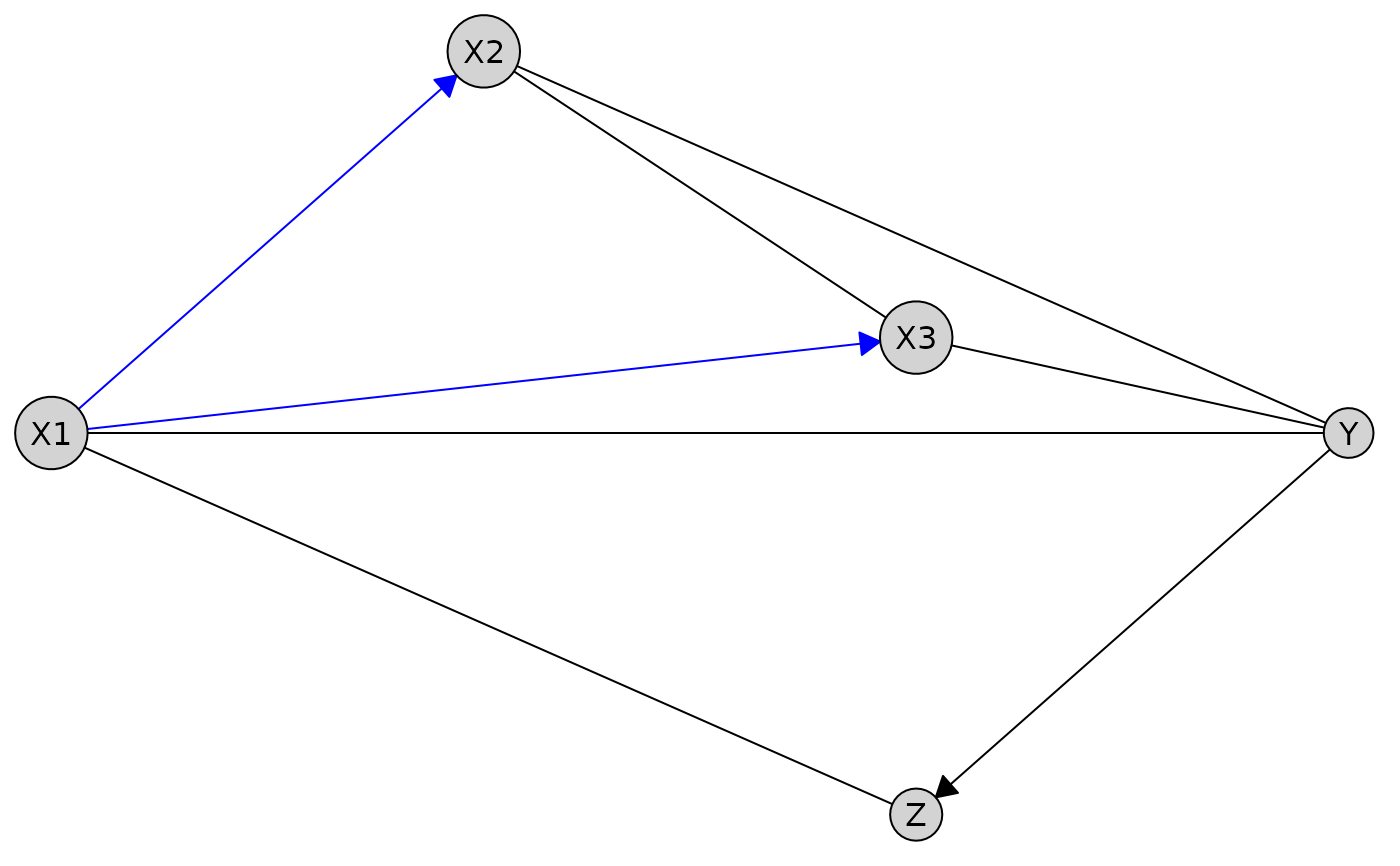
 # With a custom defined layout
custom_layout <- data.frame(
name = c("X1", "X2", "X3", "Z", "Y"),
x = c(0, 1, 2, 2, 3),
y = c(0, 1, 0.25, -1, 0)
)
plot(res_untiered, layout = custom_layout)
# With a custom defined layout
custom_layout <- data.frame(
name = c("X1", "X2", "X3", "Z", "Y"),
x = c(0, 1, 2, 2, 3),
y = c(0, 1, 0.25, -1, 0)
)
plot(res_untiered, layout = custom_layout)